Background
Overview
Teaching: 5 min
Exercises: 0 minQuestions
What is the Ocean Tracking Network?
How does your local telemetry network interact with OTN?
What methods of data analysis will be covered?
Objectives
Intro to OTN
The Ocean Tracking Network (OTN) supports global telemetry research by providing training, equipment, and data infrastructure to our large network of partners.
OTN and affiliated networks provide automated cross-referencing of your detection data with other tags in the system to help resolve “mystery detections” and provide detection data to taggers in other regions. OTN’s Data Managers will also extensively quality-control your submitted metadata for errors to ensure the most accurate records possible are stored in the database. OTN’s database and Data Portal website are excellent places to archive your datasets for future use and sharing with collaborators. We offer pathways to publish your datasets with OBIS, and via open data portals like ERDDAP, GeoServer etc. The data-product format returned by OTN is directly ingestible by analysis packages such as glatos and resonATe for ease of analysis. OTN offers support for the use of these packages and tools.
Learn more about OTN and our partners here https://members.oceantrack.org/. Please contact OTNDC@DAL.CA if you are interested in connecting with your regional network and learning about their affiliation with OTN.
Intended Audience
This set of workshop material is directed at researchers who are ready to begin the work of acoustic telemetry data analysis. The workshop begins without an introduction to the R coding language - some experience in R is expected.
If you’d like to refresh your R coding skills outside of this workshop curriculum, we recommend Data Analysis and Visualization in R for Ecologists as a good starting point. Much of this content is included in the first two lessons of this workshop.
Getting Started
Please follow the instrucions in the “Setup” tab along the top menu to install all required software, packages and data files. If you have questions or are running into errors please reach out to OTNDC@DAL.CA for support.
Intro to the pathroutr Package
The goal of pathroutr is to provide functions for re-routing paths that cross land around barrier polygons. The use-case in mind is movement paths derived from location estimates of marine animals. Due to error associated with these locations it is not uncommon for these tracks to cross land. The pathroutr package aims to provide a pragmatic and fast solution for re-routing these paths around land and along an efficient path. You can learn more here
Author: Dr. Josh M London ( josh.london@noaa.gov )
Key Points
BACKGROUND - Intro to R
Overview
Teaching: 30 min
Exercises: 20 minQuestions
What are common operators in R?
What are common data types in R?
What are some base R functions?
How do I deal with missing data?
Objectives
First, lets learn about RStudio.
RStudio is divided into 4 “Panes”: the Source for your scripts and documents (top-left, in the default layout); your Environment/History (top-right) which shows all the objects in your working space (Environment) and your command history (History); your Files/Plots/Packages/Help/Viewer (bottom-right); and the R Console (bottom-left). The placement of these panes and their content can be customized (see menu, Tools -> Global Options -> Pane Layout).
The R Script in the top pane can be saved and edited, while code typed directly into the Console below will disappear after closing the R session.
R can access files on and save outputs to any folder on your computer. R knows where to look for and save files based on the current working directory. This is one of the first things you should set up: a folder you’d like to contain all your data, scripts and outputs. The working directory path will be different for everyone. For the workshop, we’ve included the path one of our instructors uses, but you should use your computer’s file explorer to find the correct path for your data.
Setting up R
# Packages ####
# once you install packages to your computer, you can "check them out" of your packages library each time you need them
# make sure you check the "mask" messages that appear - sometimes packages have functions with the same names!
library(tidyverse)# really neat collection of packages! https://www.tidyverse.org/
library(lubridate)
library(readxl)
library(viridis)
library(plotly)
library(ggmap)
# Working Directory ####
#Instructors!! since this lesson is mostly base R we're not going to make four copies of it as with the other nodes.
#Change this one line as befits your audience.
setwd('YOUR/PATH/TO/data/NODE') #set folder you're going to work in
getwd() #check working directory
#you can also change it in the RStudio interface by navigating in the file browser where your working directory should be
#(if you can't see the folder you want, choose the three horizonal dots on the right side of the Home bar),
#and clicking on the blue gear icon "More", and select "Set As Working Directory".
Before we begin the lesson proper, a note on finding additional help. R Libraries, like those included above, are broad and contain many functions. Though most include documentation that can help if you know what to look for, sometimes more general help is necessary. To that end, RStudio maintains cheatsheets for several of the most popular libraries, which can be found here: https://www.rstudio.com/resources/cheatsheets/. As a start, the page includes an RStudio IDE cheatsheet that you may find useful while learning to navigate your workspace. With that in mind, let’s start learning R.
Intro to R
Like most programming langauges, we can do basic mathematical operations with R. These, along with variable assignment, form the basis of everything for which we will use R.
Operators
Operators in R include standard mathematical operators (+, -, *, /) as well as an assignment operator, <- (a less-than sign followed by a hyphen). The assignment operator is used to associate a value with a variable name (or, to ‘assign’ the value a name). This lets us refer to that value later, by the name we’ve given to it. This may look unfamiliar, but it fulfils the same function as the ‘=’ operator in most other languages.
3 + 5 #maths! including - , *, /
weight_kg <- 55 #assignment operator! for objects/variables. shortcut: alt + -
weight_kg
weight_lb <- 2.2 * weight_kg #can assign output to an object. can use objects to do calculations
Variables Challenge
If we change the value of weight_kg to be 100, does the value of weight_lb also change? Remember: You can check the contents of an object by typing out its name and running the line in RStudio.
Solution
No! You have to re-assign 2.2*weight_kg to the object weight_lb for it to update.
The order you run your operations is very important, if you change something you may need to re-run everything!
weight_kg <- 100 weight_lb #didnt change! weight_lb <- 2.2 * weight_kg #now its updated
Functions
While we can write code as we have in the section above - line by line, executed one line at a time - it is often more efficient to run multiple lines of code at once. By using functions, we can even compress complex calculations into just one line!
Functions use a single name to refer to underlying blocks of code that execute a specific calculation. To run a function you need two things: the name of the function, which is usually indicative of the function’s purpose; and the function’s arguments- the variables or values on which the function should execute.
#functions take "arguments": you have to tell them what to run their script against
ten <- sqrt(weight_kg) #contain calculations wrapped into one command to type.
#Output of the function can be assigned directly to a variable...
round(3.14159) #... but doesn't have to be.
Since there are hundreds of functions and often their functionality can be nuanced, we have several ways to get more information on a given function. First, we can use ‘args()’, itself a function that takes the name of another function as an argument, which will tell us the required arguments of the function against which we run it.
Second, we can use the ‘?’ operator. Typing a question mark followed by the name of a function will open a Help window in RStudio’s bottom-right panel. This will contain the most complete documentation available for the function in question.
args(round) #the args() function will show you the required arguments of another function
?round #will show you the full help page for a function, so you can see what it does
Functions Challenge
Can you round the value 3.14159 to two decimal places?
Hint: Using args() on a function can give you a clue.
Solution
round(3.14159, 2) #the round function's second argument is the number of digits you want in the result round(3.14159, digits = 2) #same as above round(digits = 2, x = 3.14159) #when reordered you need to specify
Vectors and Data Types
While variables can hold a single value, sometimes we want to store multiple values in the same variable name. For this, we can use an R data structure called a ‘vector.’ Vectors contain one or more variables of the same data type, and can be assigned to a single variable name, as below.
weight_g <- c(21, 34, 39, 54, 55) #use the combine function to join values into a vector object
length(weight_g) #explore vector
class(weight_g) #a vector can only contain one data type
str(weight_g) #find the structure of your object.
Above, we mentioned ‘data type’. This refers to the kind of data represented by a value, or stored by the appropriate variable. Data types include character (words or letters), logical (boolean TRUE or FALSE values), or numeric data. Crucially, vectors can only contain one type of data, and will force all data in the vector to conform to that type (i.e, data in the vector will all be treated as the same data type, regardless of whether or not it was of that type when the vector was created.) We can always check the data type of a variable or vector by using the ‘class()’ function, which takes the variable name as an argument.
#our first vector is numeric.
#other options include: character (words), logical (TRUE or FALSE), integer etc.
animals <- c("mouse", "rat", "dog") #to create a character vector, use quotes
class(weight_g)
class(animals)
# Note:
#R will convert (force) all values in a vector to the same data type.
#for this reason: try to keep one data type in each vector
#a data table / data frame is just multiple vectors (columns)
#this is helpful to remember when setting up your field sheets!
Vectors Challenge
What data type will this vector become?
challenge3 <- c(1, 2, 3, "4")Hint: You can check a vector’s type with the class() function.
Solution
R will force all of these to be characters, since the number 4 has quotes around it! Will always coerce data types following this structure: logical → numeric → character ← logical
class(challenge3)
Indexing and Subsetting
We can use subsetting to select only a portion of a vector. For this, we use square brackets after the name of a vector. If we supply a single numeric value, as below, we will retrieve only the value from that index of the vector. Note: vectors in R are indexed with 1 representing the first index- other languages use 0 for the start of their array, so if you are coming from a language like Python, this can be disorienting.
animals #calling your object will print it out
animals[2] #square brackets = indexing. selects the 2nd value in your vector
We can select a specific value, as above, but we can also select one or more entries based on conditions. By supplying one or more criteria to our indexing syntax, we can retrieve the elements of the array that match that criteria.
weight_g > 50 #conditional indexing: selects based on criteria
weight_g[weight_g <=30 | weight_g == 55] #many new operators here!
#<= less than or equal to; | "or"; == equal to. Also available are >=, greater than or equal to; < and > for less than or greater than (no equals); and & for "and".
weight_g[weight_g >= 30 & weight_g == 21] # >= greater than or equal to, & "and"
# this particular example give 0 results - why?
Missing Data
In practical data analysis, our data is often incomplete. It is therefore useful to cover some methods of dealing with NA values. NA is R’s shorthand for a null value; or a value where there is no data. Certain functions cannot process NA data, and therefore provide arguments that allow NA values to be removed before the function execution.
heights <- c(2, 4, 4, NA, 6)
mean(heights) #some functions cant handle NAs
mean(heights, na.rm = TRUE) #remove the NAs before calculating
This can be done within an individual function as above, but for our entire analysis we may want to produce a copy of our dataset without the NA values included. Below, we’ll explore a few ways to do that.
heights[!is.na(heights)] #select for values where its NOT NA
#[] square brackets are the base R way to select a subset of data --> called indexing
#! is an operator that reverses the function
na.omit(heights) #omit the NAs
heights[complete.cases(heights)] #select only complete cases
Missing Data Challenge
Question 1: Using the following vector of heighs in inches, create a new vector, called heights_no_na, with the NAs removed.
heights <- c(63, 69, 60, 65, NA, 68, 61, 70, 61, 59, 64, 69, 63, 63, NA, 72, 65, 64, 70, 63, 65)Solution
heights_no_na <- heights[!is.na(heights)] # or heights_no_na <- na.omit(heights) # or heights_no_na <- heights[complete.cases(heights)]Question 2: Use the function median() to calculate the median of the heights vector.
Solution
median(heights, na.rm = TRUE)Bonus question: Use R to figure out how many people in the set are taller than 67 inches.
Solution
heights_above_67 <- heights_no_na[heights_no_na > 67] length(heights_above_67)
Key Points
BACKGROUND - Starting with Data Frames
Overview
Teaching: 25 min
Exercises: 10 minQuestions
How do I import tabular data?
How do I explore my data set?
What are some basic data manipulation functions?
Objectives
Dataframes and dplyr
In this lesson, we’re going to introduce a package called dplyr. dplyr takes advantage of an operator called a pipe to create chains of data manipulation that produce powerful exploratory summaries. It also provides a suite of further functionality for manipulating dataframes: tabular sets of data that are common in data analysis. If you’ve imported the tidyverse library, as we did during setup and in the last episode, then congratulations: you already have dplyr (along with a host of other useful packages). As an aside, the cheat sheets for dplyr and readr may be useful when reviewing this lesson.
You may not be familiar with dataframes by name, but you may recognize the structure. Dataframes are arranged into rows and columns, not unlike tables in typical spreadsheet format (ex: Excel). In R, they are represented as vectors of vectors: that is, a vector wherein each column is itself a vector. If you are familiar with matrices, or two-dimensional arrays in other languages, the structure of a dataframe will be clear to you.
However, dataframes are not merely vectors- they are a specific type of object with their own functionality, which we will cover in this lesson.
We are going to use OTN-style detection extracts for this lesson. If you’re unfamiliar with detection extracts formats from OTN-style database nodes, see the documentation here
Importing from CSVs
Before we can start analyzing our data, we need to import it into R. Fortunately, we have a function for this. read_csv is a function from the readr package, also included with the tidyverse library. This function can read data from a .csv file into a dataframe. “.csv” is an extension that denotes a Comma-Separated Value file, or a file wherein data is arranged into rows, and entries within rows are delimited by commas. They’re common in data analysis.
For the purposes of this lesson, we will only cover read_csv; however, there is another function, read_excel, which you can use to import excel files. It’s from a different library (readxl) and is outside the scope of this lesson, but worth investigating if you need it.
To import your data from your CSV file, we just need to pass the file path to read_csv, and assign the output to a variable. Note that the file path you give to read_csv will be relative to the working directory you set in the last lesson, so keep that in mind.
#imports file into R. paste the filepath to the file here!
#read_csv can take both csv and zip files, as long as the zip file contains a csv.
nsbs_matched_2021 <- read_csv("nsbs_matched_detections_2021.zip")
We can now refer to the variable nsbs_matched_2021 to access, manipulate, and view the data from our CSV. In the next sections, we will explore some of the basic operations you can perform on dataframes.
Exploring Detection Extracts
Let’s start with a practical example. What can we find out about these matched detections? We’ll begin by running the code below, and then give some insight into what each function does. Remember, if you’re ever confused about the purpose of a function, you can use ‘?’ followed by the function name (i.e, ?head, ?View) to get more information.
head(nsbs_matched_2021) #first 6 rows
View(nsbs_matched_2021) #can also click on object in Environment window
str(nsbs_matched_2021) #can see the type of each column (vector)
glimpse(nsbs_matched_2021) #similar to str()
#summary() is a base R function that will spit out some quick stats about a vector (column)
#the $ syntax is the way base R selects columns from a data frame
summary(nsbs_matched_2021$latitude)
You may now have an idea of what each of those functions does, but we will briefly explain each here.
head takes the dataframe as a parameter and returns the first 6 rows of the dataframe. This is useful if you want to quickly check that a dataframe imported, or that all the columns are there, or see other such at-a-glance information. Its primary utility is that it is much faster to load and review than the entire dataframe, which may be several tens of thousands of rows long. Note that the related function tail will return the last six elements.
If we do want to load the entire dataframe, though, we can use View, which will open the dataframe in its own panel, where we can scroll through it as though it were an Excel file. This is useful for seeking out specific information without having to consult the file itself. Note that for large dataframes, this function can take a long time to execute.
Next are the functions str and glimpse, which do similar but different things. str is short for ‘structure’ and will print out a lot of information about your dataframe, including the number of rows and columns (called ‘observations’ and ‘variables’), the column names, the first four entries of each column, and each column type as well. str can sometimes be a bit overwhelming, so if you want to see a more condensed output, glimpse can be useful. It prints less information, but is cleaner and more compact, which can be desirable.
Finally, we have the summary function, which takes a single column from a dataframe and produces a summary of its basic statistics. You may have noticed the ‘$’ in the summary call- this is how we index a specific column from a dataframe. In this case, we are referring to the latitude column of our dataframe.
Using what you now know about summary functions, try to answer the challenge below.
Detection Extracts Challenge
Question 1: What is the class of the station column in nsbs_matched_2021, and how many rows and columns are in the nsbs_matched_2021 dataset??
Solution
The column is a character, and there are 7,693 rows with 36 columns
str(nsbs_matched_2021) # or glimpse(nsbs_matched_2021)
Data Manipulation
Now that we’ve learned how to import and summarize our data, we can learn how to use dplyr to manipulate it. The name ‘dplyr’ may seem esoteric- the ‘d’ is short for ‘dataframe’, and ‘plyr’ is meant to evoke pliers, and thereby cutting, twisting, and shaping. This is an elegant summation of the dplyr library’s functionality.
We are going to introduce a new operator in this section, called the “dplyr pipe”. Not to be confused with |, which is also called a pipe in some other languages, the dplyr pipe is rendered as %>%. A pipe takes the output of the function or contents of the variable on the left and passes them to the function on the right. It is often read as “and then.” If you want to quickly add a pipe, the keybord shortcut CTRL + SHIFT + M will do so.
library(dplyr) #can use tidyverse package dplyr to do exploration on dataframes in a nicer way
# %>% is a "pipe" which allows you to join functions together in sequence.
nsbs_matched_2021 %>% dplyr::select(6) #selects column 6
# Using the above transliteration: "take nsbs_matched_2021 AND THEN select column number 6 from it using the select function in the dplyr library"
You may have noticed another unfamiliar operator above, the double colon (::). This is used to specify the package from which we want to pull a function. Until now, we haven’t needed this, but as your code grows and the number of libraries you’re using increases, it’s likely that multiple functions across several different packages will have the same name (a phenomenon called “overloading”). R has no automatic way of knowing which package contains the function you are referring to, so using double colons lets us specify it explicitly. It’s important to be able to do this, since different functions with the same name often do markedly different things.
Let’s explore a few other examples of how we can use dplyr and pipes to manipulate our dataframe.
nsbs_matched_2021 %>% slice(1:5) #selects rows 1 to 5 in the dplyr way
# Take nsbs_matched_2021 AND THEN slice rows 1 through 5.
#We can also use multiple pipes.
nsbs_matched_2021 %>%
distinct(detectedby) %>% nrow #number of arrays that detected my fish in dplyr!
# Take nsbs_matched_2021 AND THEN select only the unique entries in the detectedby column AND THEN count them with nrow.
#We can do the same as above with other columns too.
nsbs_matched_2021 %>%
distinct(catalognumber) %>%
nrow #number of animals that were detected
# Take nsbs_matched_2021 AND THEN select only the unique entries in the catalognumber column AND THEN count them with nrow.
#We can use filtering to conditionally select rows as well.
nsbs_matched_2021 %>% filter(catalognumber=="NSBS-1393332-2021-08-05")
# Take nsbs_matched_2021 AND THEN select only those rows where catalognumber is equal to the above value.
nsbs_matched_2021 %>% filter(monthcollected >= 10) #all dets in/after October of 2016
# Take nsbs_matched_2021 AND THEN select only those rows where monthcollected is greater than or equal to 10.
These are all ways to extract a specific subset of our data, but dplyr can also be used to manipulate dataframes to give you even greater insights. We’re now going to use two new functions: group_by, which allows us to group our data by the values of a single column, and summarise (not to be confused with summary above!), which can be used to calculate summary statistics across your grouped variables, and produces a new dataframe containing these values as the output. These functions can be difficult to grasp, so don’t forget to use ?group_by and ?summarise if you get lost.
#get the mean value across a column using GroupBy and Summarize
nsbs_matched_2021 %>% #Take nsbs_matched_2021, AND THEN...
group_by(catalognumber) %>% #Group the data by catalognumber- that is, create a group within the dataframe where each group contains all the rows related to a specific catalognumber. AND THEN...
summarise(MeanLat=mean(latitude)) #use summarise to add a new column containing the mean latitude of each group. We named this new column "MeanLat" but you could name it anything
With just a few lines of code, we’ve created a dataframe that contains each of our catalog numbers and the mean latitude at which those fish were detected. dplyr, its wide array of functions, and the powerful pipe operator can let us build out detailed summaries like this one without writing too much code.
Data Manipulation Challenge
Question 1: Find the max lat and max longitude for animal “NSBS-1393332-2021-08-05”.
Solution
nsbs_matched_2021 %>% filter(catalognumber=="NSBS-1393332-2021-08-05") %>% summarise(MaxLat=max(latitude), MaxLong=max(longitude))Question 2: Find the min lat/long of each animal for detections occurring in/after April.
Solution
nsbs_matched_2021 %>% filter(monthcollected >= 4 ) %>% group_by(catalognumber) %>% summarise(MinLat=min(latitude), MinLong=min(longitude))
Joining Detection Extracts
We’re now going to briefly touch on a few useful dataframe use-cases that aren’t directly related to dplyr, but with which dplyr can help us.
One function that we’ll need to know is rbind, a base R function which lets us combine two R objects together. Since detections for animals tagged during a study often appear in multiple years, this functionality will let us merge the dataframes together. We’ll also use distinct, a dplyr function that lets us trim out duplicate release records for each animal, since these are listed in each detection extract.
nsbs_matched_2022 <- read_csv("nsbs_matched_detections_2022.zip") #First, read in our file.
nsbs_matched_full <- rbind(nsbs_matched_2021, nsbs_matched_2022) #Now join the two dataframes
# release records for animals often appear in >1 year, this will remove the duplicates
nsbs_matched_full <- nsbs_matched_full %>% distinct() # Use distinct to remove duplicates.
View(nsbs_matched_full)
Dealing with Datetimes
Datetime data is in a special format which is neither numeric nor character. It can be tricky to deal with, too, since Excel frequently reformats dates in any file it opens. We also have to concern ourselves with practical matters of time, like time zone and date formatting. Fortunately, the lubridate library gives us a whole host of functionality to manage datetime data. For additional help, the cheat sheet for lubridate may prove a useful resource.
We’ll also use a dplyr function called mutate, which lets us add new columns or change existing ones, while preserving the existing data in the table. Be careful not to confuse this with its sister function transmute, which adds or manipulates columns while dropping existing data. If you’re ever in doubt as to which is which, remember: ?mutate and ?transmute will bring up the help files.
library(lubridate) #Import our Lubridate library.
nsbs_matched_full %>% mutate(datecollected=ymd_hms(datecollected)) #Use the lubridate function ymd_hms to change the format of the date.
#as.POSIXct(nsbs_matched_full$datecollected) #this is the base R way - if you ever see this function
We’ve just used a single function, ymd_hms, but with it we’ve been able to completely reformat the entire datecollected column. ymd_hms is short for Year, Month, Day, Hours, Minutes, and Seconds. For example, at time of writing, it’s 2021-05-14 14:21:40. Other format functions exist too, like dmy_hms, which specifies the day first and year third (i.e, 14-05-2021 14:21:40). Investigate the documentation to find which is right for you.
There are too many useful lubridate functions to cover in the scope of this lesson. These include parse_date_time, which can be used to read in date data in multiple formats, which is useful if you have a column contianing heterogenous date data; as well as with_tz, which lets you make your data sensitive to timezones (including automatic daylight savings time awareness). Dates are a tricky subject, so be sure to investigate lubridate to make sure you find the functions you need.
Key Points
BACKGROUND - Intro to Plotting
Overview
Teaching: 15 min
Exercises: 10 minQuestions
How do I plot my data?
How can I plot summaries of my data?
Objectives
Learn how to make basic plots with ggplot2
Learn how to combine dplyr summaries with ggplot2 plots
Background
Now that we have learned how to import, inspect, and manipulate our data, we are next going to learn how to visualize it. R provides a robust plotting suite in the library ggplot2. ggplot2 takes advantage of tidyverse pipes and chains of data manipulation to build plotting code. Additionally, it separates the aesthetics of the plot (what are we plotting) from the styling of the plot (what the plot looks like). What this means is that data aesthetics and styles can be built separately and then combined and recombined to produce modular, reusable plotting code. If ggplot seems daunting, the cheat sheet may prove useful.
While ggplot2 function calls can look daunting at first, they follow a single formula, detailed below.
#Anything within <> braces will be replaced in an actual function call.
ggplot(data = <DATA>, mapping = aes(<MAPPINGS>)) + <GEOM_FUNCTION>
In the above example, there are three important parts: <DATA>, <MAPPINGS>, and <GEOM_FUNCTION>.
<DATA> refers to the data that we’ll be plotting. In general, this will be held in a dataframe like the one we prepared in the previous lessons.
<MAPPINGS> refers to the aesthetic mappings for the data- that is, which columns in the data will be used to determine which attributes of the graph. For example, if you have columns for latitude and longitude, you may want to map these onto the X and Y axes of the graph. We’ll cover how to do exactly that in a moment.
Finally, <GEOM_FUNCTION> refers to the style of the plot: what type of plot are we going to make. GEOM is short for “geometry” and ggplot2 contains many different ‘geom’ functions that you can use. For this lesson, we’ll be using geom_point(), which produces a scatterplot, but in the future you may want to use geom_path(), geom_bar(), geom_boxplot() or any of ggplots other geom functions. Remember, since these are functions, you can use the help syntax (i.e ?geom_point) in the R console to find out more about them and what you need to pass to them.
Now that we’ve introduced ggplot2, let’s build a functional example with our data.
# Begin by importing the ggplot2 library, which you should have installed as part of setup.
library(ggplot2)
# Build the plot and assign it to a variable.
nsbs_matched_full_plot <- ggplot(data = nsbs_matched_full,
mapping = aes(x = longitude, y = latitude)) #can assign a base
With a couple of lines of code, we’ve already mostly completed a simple scatter plot of our data. The ‘data’ parameter takes our dataframe, and the mapping parameter takes the output of the aes() function, which itself takes a mapping of our data onto the axes of the graph. That can be a bit confusing, so let’s briefly break this down. aes() is short for ‘aesthetics’- the function constructs the aesthetic mappings of our data, which describe how variables in the data are mapped to visual properties of the plot. For example, above, we are setting the ‘x’ attribute to ‘longitude’, and the ‘y’ attribute to latitude. This means that the X axis of our plot will represent longitude, and the Y axis will represent latitude. Depending on the type of plot you’re making, you may want different values there, and different types of geom functions can require different aesthetic mappings (colour, for example, is another common one). You can always type ?aes() at the console if you want more information.
We still have one step to add to our plotting code: the geom function. We’ll be making a scatterplot, so we want to use geom_point().
nsbs_matched_full_plot +
geom_point(alpha=0.1,
colour = "blue")
#This will layer our chosen geom onto our plot template.
#alpha is a transparency argument in case points overlap. Try alpha = 0.02 to see how it works!
With just the above code, we’ve added our geom to our aesthetic and made our plot ready for display. We’ve built only a very simple plot here, but ggplot2 provides many, many options for building more complex, illustrative plots.
Basic plots
As a minor syntactic note, you can build your plots iteratively, without assigning them to a variable in-between. For this, we make use of tidyverse pipes.
nsbs_matched_full %>%
ggplot(aes(longitude, latitude)) +
geom_point() #geom = the type of plot
nsbs_matched_full %>%
ggplot(aes(longitude, latitude, colour = commonname)) +
geom_point()
#anything you specify in the aes() is applied to the actual data points/whole plot,
#anything specified in geom() is applied to that layer only (colour, size...). sometimes you have >1 geom layer so this makes more sense!
You can see that all we need to do to make this work is omit the ‘data’ parameter, since that’s being passed in by the pipe. Note also that we’ve added colour = commonname to the second plot’s aesthetic, meaning that the output will be coloured based on the species of the animal (if there is more than one included).
Remembering which of the aes or the geom controls which variable can be difficult, but here’s a handy rule of thumb: anything specified in aes() will apply to the data points themselves, or the whole plot. They are broad statements about how the plot is to be displayed. Anything in the geom_ function will apply only to that geom_ layer. Keep this in mind, since it’s possible for your plot to have more than one geom_!
Plotting and dplyr Challenge
Try combining with
dplyrfunctions in this challenge! Try making a scatterplot showing the lat/long for animal “NSBS-1393332-2021-08-05”, coloured by detection arraySolution
nsbs_matched_full %>% filter(catalognumber=="NSBS-1393332-2021-08-05") %>% ggplot(aes(longitude, latitude, colour = detectedby)) + geom_point()What other geoms are there? Try typing
geom_into R to see what it suggests!
Key Points
You can feed output from dplyr’s data manipulation functions into ggplot using pipes.
Plotting various summaries and groupings of your data is good practice at the exploratory phase, and dplyr and ggplot make iterating different ideas straightforward.
Telemetry Reports - Imports
Overview
Teaching: 10 min
Exercises: 0 minQuestions
What datasets do I need from the Network?
How do I import all the datasets?
Objectives
Importing all the datasets
Let’s look at how we might implement a common telemetry workflow using Tidyverse libraries like dplyr and ggplot2.
We are going to use OTN-style detection extracts for this lesson. If you’re unfamiliar with detection extracts formats from OTN-style database nodes, see the documentation here.
For OTN you will receive Detection Extracts which include (1) Matched to Animals YYYY, (2) Detections Mapped to Other Trackers YYYY (also called Qualified) and (3) Unqualified Detections YYYY. In each case, the YYYY in the filename indicates the single year of data contained in the file. The types of detection extracts you receive will differ depending on the type of project you have regitered with the Network. ex: Tag-only projects will not receive Qualified and Unqualified detection extracts.
To illustrate the many meaningful summary reports which can be created use detection extracts, we will import an example of Matched and Qualified extracts.
First, we will comfirm we have our Tag Matches stored in a dataframe.
library(tidyverse)# really neat collection of packages! https://www.tidyverse.org/
library(lubridate)
library(readxl)
library(viridis)
library(plotly)
library(ggmap)
setwd('YOUR/PATH/TO/data/otn') #set folder you're going to work in
getwd() #check working directory
nsbs_matched_2021 <- read_csv("nsbs_matched_detections_2021.zip") #Import 2021 detections
nsbs_matched_2022 <- read_csv("nsbs_matched_detections_2022.zip") # Import 2022 detections
nsbs_matched_full <- rbind(nsbs_matched_2021, nsbs_matched_2022) #Now join the two dataframes
# release records for animals often appear in >1 year, this will remove the duplicates
nsbs_matched_full <- nsbs_matched_full %>% distinct() # Use distinct to remove duplicates.
Next, we will load in and join our Array matches. Ensure you replace the filepath to show the files as they appear in your working directory, if needed.
hfx_qual_2021 <- read_csv("hfx_qualified_detections_2021_workshop.csv")
hfx_qual_2022 <- read_csv("hfx_qualified_detections_2022_workshop.csv")
hfx_qual_21_22_full <- rbind(hfx_qual_2021, hfx_qual_2022)
To give meaning to these detections we should import our Instrument Deployment Metadata and Tagging Metadata as well. These are in the standard OTN-style templates which can be found here.
#These are saved as XLS/XLSX files, so we need a different library to read them in.
# Deployment Metadata
hfx_deploy <- read_excel("hfx_sample_deploy_metadata_export.xlsx", skip=3) #can also do argument "sheet = XXX" if needed
View(hfx_deploy)
# Tag metadata
nsbs_tag <- read_excel("nsbs_sample_tag_metadata_export.xlsx")
View(nsbs_tag)
#keep in mind the timezone of the columns
Key Points
Telemetry Reports for Array Operators
Overview
Teaching: 30 min
Exercises: 0 minQuestions
How do I summarize and plot my deployments?
How do I summarize and plot my detections?
Objectives
Mapping our stations - Static map
We can do the same exact thing with the deployment metadata from OUR project only!
names(hfx_deploy)
base <- get_stadiamap(
bbox = c(left = min(hfx_deploy$DEPLOY_LONG),
bottom = min(hfx_deploy$DEPLOY_LAT),
right = max(hfx_deploy$DEPLOY_LONG),
top = max(hfx_deploy$DEPLOY_LAT)),
maptype = "stamen_toner_lite",
crop = FALSE,
zoom = 5)
#filter for stations you want to plot - this is very customizable
hfx_deploy_plot <- hfx_deploy %>%
mutate(deploy_date=ymd_hms(`DEPLOY_DATE_TIME (yyyy-mm-ddThh:mm:ss)`)) %>% #make a datetime
mutate(recover_date=ymd_hms(`RECOVER_DATE_TIME (yyyy-mm-ddThh:mm:ss)`)) %>% #make a datetime
filter(!is.na(deploy_date)) %>% #no null deploys
filter(deploy_date > '2020-07-03' | recover_date < '2022-01-11') %>% #only looking at certain deployments, can add start/end dates here
group_by(STATION_NO) %>%
summarise(MeanLat=mean(DEPLOY_LAT), MeanLong=mean(DEPLOY_LONG)) #get the mean location per station, in case there is >1 deployment
#add your stations onto your basemap
hfx_map <-
ggmap(base, extent='panel') +
ylab("Latitude") +
xlab("Longitude") +
geom_point(data = hfx_deploy_plot, #filtering for recent deployments
aes(x = MeanLong,y = MeanLat), #specify the data, colour = STATION_NO is also neat here
shape = 19, size = 2) #lots of aesthetic options here!
#view your receiver map!
hfx_map
#save your receiver map into your working directory
ggsave(plot = hfx_map, filename = "hfx_map.tiff", units="in", width=15, height=8)
#can specify location, file type and dimensions
Mapping my stations - Interactive map
An interactive map can contain more information than a static map. Here we will explore the package plotly to create interactive “slippy” maps. These allow you to explore your map in different ways by clicking and scrolling through the output.
First, we will set our basemap’s aesthetics and bounding box and assign this information (as a list) to a geo_styling variable.
library(plotly)
#set your basemap
geo_styling <- list(
scope = 'nova scotia',
fitbounds = "locations", visible = TRUE, #fits the bounds to your data!
showland = TRUE,
showlakes = TRUE,
lakecolor = toRGB("blue", alpha = 0.2), #make it transparent
showcountries = TRUE,
landcolor = toRGB("gray95"),
countrycolor = toRGB("gray85")
)
Then, we choose which Deployment Metadata dataset we wish to use and identify the columns containing Latitude and Longitude, using the plot_geo function.
#decide what data you're going to use. Let's use hfx_deploy_plot, which we created above for our static map.
hfx_map_plotly <- plot_geo(hfx_deploy_plot, lat = ~MeanLat, lon = ~MeanLong)
Next, we use the add_markers function to write out what information we would like to have displayed when we hover our mouse over a station in our interactive map. In this case, we chose to use paste to join together the Station Name and its lat/long.
#add your markers for the interactive map
hfx_map_plotly <- hfx_map_plotly %>% add_markers(
text = ~paste(STATION_NO, MeanLat, MeanLong, sep = "<br />"),
symbol = I("square"), size = I(8), hoverinfo = "text"
)
Finally, we add all this information together, along with a title, using the layout function, and now we can explore our interactive map!
#Add layout (title + geo stying)
hfx_map_plotly <- hfx_map_plotly %>% layout(
title = 'HFX Deployments<br />(> 2020-07-03)', geo = geo_styling
)
#View map
hfx_map_plotly
To save this interactive map as an .html file, you can explore the function htmlwidgets::saveWidget(), which is beyond the scope of this lesson.
Summary of Animals Detected
Let’s find out more about the animals detected by our array! These summary statistics, created using dplyr functions, could be used to help determine the how successful each of your stations has been at detecting tagged animals. We will also learn how to export our results using write_csv.
# How many of each animal did we detect from each collaborator, per station
library(dplyr)
hfx_qual_summary <- hfx_qual_21_22_full %>%
filter(datecollected > '2021-06-01') %>% #select timeframe, stations etc.
group_by(trackercode, station, tag_contact_pi, tag_contact_poc) %>%
summarize(count = n()) %>%
dplyr::select(trackercode, tag_contact_pi, tag_contact_poc, station, count)
#view our summary table
view(hfx_qual_summary)
#export our summary table
write_csv(hfx_qual_summary, "hfx_summary.csv", col_names = TRUE)
Summary of Detections
These dplyr summaries can suggest array performance, hotspot stations, and be used as a metric for funders.
# number of detections per month/year per station
hfx_det_summary <- hfx_qual_21_22_full %>%
mutate(datecollected=ymd_hms(datecollected)) %>%
group_by(station, year = year(datecollected), month = month(datecollected)) %>%
summarize(count =n())
hfx_det_summary
# Create a new data product, det_days, that give you the unique dates that an animal was seen by a station
stationsum <- hfx_qual_21_22_full %>%
group_by(station) %>%
summarise(num_detections = length(datecollected),
start = min(datecollected),
end = max(datecollected),
uniqueIDs = length(unique(fieldnumber)),
det_days=length(unique(as.Date(datecollected))))
View(stationsum)
Plot of Detections
Lets make an informative plot using ggplot showing the number of matched detections, per year and month. Remember: we can combine dplyr data manipulation and plotting into one step, using pipes!
hfx_qual_21_22_full %>%
mutate(datecollected=ymd_hms(datecollected)) %>% #make datetime
mutate(year_month = floor_date(datecollected, "months")) %>% #round to month
group_by(year_month) %>% #can group by station, species etc.
summarize(count =n()) %>% #how many dets per year_month
ggplot(aes(x = (month(year_month) %>% as.factor()),
y = count,
fill = (year(year_month) %>% as.factor())
)
)+
geom_bar(stat = "identity", position = "dodge2")+
xlab("Month")+
ylab("Total Detection Count")+
ggtitle('HFX Animal Detections by Month')+ #title
labs(fill = "Year") #legend title
Key Points
Telemetry Reports for Tag Owners
Overview
Teaching: 30 min
Exercises: 0 minQuestions
How do I summarize and plot my detections?
How do I summarize and plot my tag metadata?
Objectives
New dataframes
To aid in the creating of useful Matched Detection summaries, we should create a new dataframe where we filter out release records from the detection extracts. This will leave only “true” detections.
#optional dataset to use: detections with releases filtered out!
nsbs_matched_full_no_release <- nsbs_matched_full %>%
filter(receiver != "release")
Mapping my Detections and Releases - static map
Where were my fish observed? We will make a static map of all the receiver stations where my fish was detected in two steps, using the package ggmap.
First, we set a basemap using the aesthetics and bounding box we desire. Next, we add the detection locations onto the basemap and look at our creation!
base <- get_stadiamap(
bbox = c(left = min(nsbs_matched_full_no_release$longitude),
bottom = min(nsbs_matched_full_no_release$latitude),
right = max(nsbs_matched_full_no_release$longitude),
top = max(nsbs_matched_full_no_release$latitude)),
maptype = "stamen_toner_lite",
crop = FALSE,
zoom = 5)
#add your releases and detections onto your basemap
nsbs_map <-
ggmap(base, extent='panel') +
ylab("Latitude") +
xlab("Longitude") +
geom_point(data = nsbs_matched_full_no_release,
aes(x = longitude,y = latitude), #specify the data
colour = 'blue', shape = 19, size = 2) #lots of aesthetic options here!
#view your tagging map!
nsbs_map
Mapping my Detections and Releases - interactive map
An interactive map can contain more information than a static map. Here we will explore the package plotly to create interactive “slippy” maps. These allow you to explore your map in different ways by clicking and scrolling through the output.
First, we will set our basemap’s aesthetics and bounding box and assign this information (as a list) to a geo_styling variable. Then, we choose which detections we wish to use and identify the columns containing Latitude and Longitude, using the plot_geo function. Next, we use the add_markers function to write out what information we would like to have displayed when we hover our mouse over a station in our interactive map. In this case, we chose to use paste to join together the Station Name and its lat/long. Finally, we add all this information together, along with a title, using the layout function, and now we can explore our interactive map!
#set your basemap
geo_styling <- list(
fitbounds = "locations", visible = TRUE, #fits the bounds to your data!
showland = TRUE,
showlakes = TRUE,
lakecolor = toRGB("blue", alpha = 0.2), #make it transparent
showcountries = TRUE,
landcolor = toRGB("gray95"),
countrycolor = toRGB("gray85")
)
#decide what data you're going to use
detections_map_plotly <- plot_geo(nsbs_matched_full_no_release, lat = ~latitude, lon = ~longitude)
#add your markers for the interactive map
detections_map_plotly <- detections_map_plotly %>% add_markers(
text = ~paste(catalognumber, commonname, paste("Date detected:", datecollected),
paste("Latitude:", latitude), paste("Longitude",longitude),
paste("Detected by:", detectedby), paste("Station:", station),
paste("Project:",collectioncode), sep = "<br />"),
symbol = I("square"), size = I(8), hoverinfo = "text"
)
#Add layout (title + geo stying)
detections_map_plotly <- detections_map_plotly %>% layout(
title = 'NSBS Detections', geo = geo_styling
)
#View map
detections_map_plotly
Summary of tagged animals
This section will use your Tagging Metadata to create dplyr summaries of your tagged animals.
# summary of animals you've tagged
nsbs_tag_summary <- nsbs_tag %>%
mutate(UTC_RELEASE_DATE_TIME = ymd_hms(UTC_RELEASE_DATE_TIME)) %>%
#filter(UTC_RELEASE_DATE_TIME > '2016-06-01') %>% #select timeframe, specific animals etc.
group_by(year = year(UTC_RELEASE_DATE_TIME), COMMON_NAME_E) %>%
summarize(count = n(),
Meanlength = mean(`LENGTH (m)`, na.rm=TRUE),
minlength= min(`LENGTH (m)`, na.rm=TRUE),
maxlength = max(`LENGTH (m)`, na.rm=TRUE),
MeanWeight = mean(`WEIGHT (kg)`, na.rm = TRUE))
#view our summary table
View(nsbs_tag_summary)
Detection Attributes
Lets add some biological context to our summaries! To do this we can join our Tag Metadata with our Matched Detections. To learn more about the different types of dataframe joins and how they function, see here.
# Average location of each animal, without release records
nsbs_matched_full_no_release %>%
group_by(catalognumber) %>%
summarize(NumberOfStations = n_distinct(station),
AvgLat = mean(latitude),
AvgLong =mean(longitude))
Now lets try to join our metadata and detection extracts.
#First we need to make a tagname column in the tag metadata (to match the Detection Extract), and figure out the enddate of the tag battery.
nsbs_tag <- nsbs_tag %>%
mutate(enddatetime = (ymd_hms(UTC_RELEASE_DATE_TIME) + days(EST_TAG_LIFE))) %>% #adding enddate
mutate(tagname = paste(TAG_CODE_SPACE,TAG_ID_CODE, sep = '-')) #adding tagname column
#Now we join by tagname, to the detection dataset (without the release information)
tag_joined_dets <- left_join(x = nsbs_matched_full_no_release, y = nsbs_tag, by = "tagname") #join!
#make sure any redeployed tags have matched within their deployment period only
tag_joined_dets <- tag_joined_dets %>%
filter(datecollected >= UTC_RELEASE_DATE_TIME & datecollected <= enddatetime)
View(tag_joined_dets)
Lets use this new joined dataframe to make summaries!
#Avg length per location
nsbs_tag_det_summary <- tag_joined_dets %>%
group_by(detectedby, station, latitude, longitude) %>%
summarise(AvgSize = mean(`LENGTH (m)`, na.rm=TRUE))
View(nsbs_tag_det_summary)
#export our summary table as CSV
write_csv(nsbs_tag_det_summary, "detections_summary.csv", col_names = TRUE)
# count detections per transmitter, per array
nsbs_matched_full_no_release %>%
group_by(catalognumber, station, detectedby, commonname) %>%
summarize(count = n()) %>%
dplyr::select(catalognumber, commonname, detectedby, station, count)
# list all receivers each fish was seen on, and a number_of_receivers column too
arrays <- nsbs_matched_full_no_release %>%
group_by(catalognumber) %>%
mutate(arrays = (list(unique(detectedby)))) %>% #create a column with a list of the stations
dplyr::select(catalognumber, arrays) %>% #remove excess columns
distinct_all() %>% #keep only one record of each
mutate(number_of_arrays = sapply(arrays, length)) %>% #sapply: applies a function across a List - in this case we are applying length()
as.data.frame()
View(arrays)
# number of stations visited, start and end dates, and track length
animal_id_summary <- nsbs_matched_full_no_release %>%
group_by(catalognumber) %>%
summarise(dets = length(catalognumber),
stations = length(unique(station)),
min = min(datecollected),
max = max(datecollected),
tracklength = max(datecollected)-min(datecollected))
View(animal_id_summary)
Summary of Detection Counts
Lets make an informative plot showing number of matched detections, per year and month.
nsbs_matched_full_no_release %>%
mutate(datecollected=ymd_hms(datecollected)) %>% #make datetime
mutate(year_month = floor_date(datecollected, "months")) %>% #round to month
group_by(year_month) %>% #can group by station, species etc.
summarize(count =n()) %>% #how many dets per year_month
ggplot(aes(x = (month(year_month) %>% as.factor()),
y = count,
fill = (year(year_month) %>% as.factor())
)
)+
geom_bar(stat = "identity", position = "dodge2")+
xlab("Month")+
ylab("Total Detection Count")+
ggtitle('NSBS Detections by Month (2021-2022)')+ #title
labs(fill = "Year") #legend title
Other Example Plots
Some examples of complex plotting options. The most useful of these may include abacus plotting (an example with ‘animal’ and ‘station’ on the y-axis) as well as an example using ggmap and geom_path to create an example map showing animal movement.
#Use the color scales in this package to make plots that are pretty,
#better represent your data, easier to read by those with colorblindness, and print well in grey scale.
library(viridis)
# an easy abacus plot!
abacus_animals <-
ggplot(data = nsbs_matched_full, aes(x = datecollected, y = catalognumber, col = detectedby)) +
geom_point() +
ggtitle("Detections by animal") +
theme(plot.title = element_text(face = "bold", hjust = 0.5)) +
scale_color_viridis(discrete = TRUE)
abacus_animals
abacus_arrays <-
ggplot(data = nsbs_matched_full, aes(x = datecollected, y = detectedby, col = catalognumber)) +
geom_point() +
ggtitle("Detections by Array") +
theme(plot.title = element_text(face = "bold", hjust = 0.5)) +
scale_color_viridis(discrete = TRUE)
abacus_arrays #might be better with just a subset, huh??
# track movement using geom_path!!
nsbs_subset <- nsbs_matched_full %>%
dplyr::filter(catalognumber %in% c('NSBS-Nessie', 'NSBS-1250981-2019-09-06',
'NSBS-1393342-2021-08-10', 'NSBS-1393332-2021-08-05'))
View(nsbs_subset)
movMap <-
ggmap(base, extent = 'panel') + #use the BASE we set up before
ylab("Latitude") +
xlab("Longitude") +
geom_path(data = nsbs_subset, aes(x = longitude, y = latitude, col = commonname)) + #connect the dots with lines
geom_point(data = nsbs_subset, aes(x = longitude, y = latitude, col = commonname)) + #layer the stations back on
#scale_colour_manual(values = c("red", "blue"), name = "Species")+ #for more than one species
facet_wrap(~catalognumber, nrow=2, ncol=2)+
ggtitle("Inferred Animal Paths")
#to size the dots by number of detections you could do something like: size = (log(length(animal)id))?
movMap
# monthly latitudinal distribution of your animals (works best w >1 species)
nsbs_matched_full %>%
group_by(m=month(datecollected), catalognumber, scientificname) %>% #make our groups
summarise(mean=mean(latitude)) %>% #mean lat
ggplot(aes(m %>% factor, mean, colour=scientificname, fill=scientificname))+ #the data is supplied, but no info on how to show it!
geom_point(size=3, position="jitter")+ # draw data as points, and use jitter to help see all points instead of superimposition
#coord_flip()+ #flip x y, not needed here
scale_colour_manual(values = "blue")+ #change the colour to represent the species better!
scale_fill_manual(values = "grey")+
geom_boxplot()+ #another layer
geom_violin(colour="black") #and one more layer
#There are other ways to present a summary of data like this that we might have chosen.
#geom_density2d() will give us a KDE for our data points and give us some contours across our chosen plot axes.
nsbs_matched_full %>%
group_by(month=month(datecollected), catalognumber, scientificname) %>%
summarise(meanlat=mean(latitude)) %>%
ggplot(aes(month, meanlat, colour=scientificname, fill=scientificname))+
geom_point(size=3, position="jitter")+
scale_colour_manual(values = "blue")+
scale_fill_manual(values = "grey")+
geom_density2d(linewidth=7, lty=1) #this is the only difference from the plot above
#anything you specify in the aes() is applied to the actual data points/whole plot,
#anything specified in geom() is applied to that layer only (colour, size...)
# per-individual density contours - lots of plots: called facets!
nsbs_matched_full %>%
ggplot(aes(longitude, latitude))+
facet_wrap(~catalognumber)+ #make one plot per individual
geom_violin()
#Warnings going on above.
Key Points
Basic Animation
Overview
Teaching: 20 min
Exercises: 0 minQuestions
How do I set up my data extract for animation?
How do I animate my animal movements?
Objectives
Static plots are excellent tools and are appropriate a lot of the time, but there are instances where something extra is needed to demonstrate your interesting fish movements. This is where plotting animated tracks can be a useful tool. In this lesson we will explore how to take data from your OTN-style detection extract documents and animate the journey of one fish between stations.
Getting our Packages
If not done already, we will first need to ensure we have all the required packages activated in our R session.
library(glatos)
library(sf)
library(mapview)
library(plotly)
library(gganimate)
library(ggmap)
library(tidyverse)
Preparing our Dataset
Before we can animate, we need to do some preprocessing on our dataset. For this animation we will be using detection events (a format we learned about in the glatos lessons) so we will need to first create that variable. To do this, we will read in our data using the read_otn_detections function from glatos and check for false detections with the false_detections function.
For the purposes of this lesson we will assume that any detection that did not pass the filter is a false detection, and will filter them out using filter(passed_filter != FALSE). It is important to note that for real data you will need to look over these detections to be sure they are truly false.
Finally, we use the detection_events function with station as the location_col argument to get our detection events.
unzip('nsbs_matched_detections_2022.zip', overwrite = TRUE)
detection_events <- #create detections event variable
read_otn_detections('nsbs_matched_detections_2022/nsbs_matched_detections_2022.csv') %>%
false_detections(tf = 3600) %>% #find false detections
dplyr::filter(passed_filter != FALSE) %>%
detection_events(location_col = 'station', time_sep=3600)
There is extra information in detection_events (such as the number of detections per event and the residence time in seconds) that can make some interesting plots, but for our visualization we only need the animal_id, mean_longitude, mean_latitude, and first_detection columns. So we will use the dplyr select function to create a dataframe with just those columns.
plot_data <- detection_events %>%
dplyr::select(animal_id, mean_longitude,mean_latitude, first_detection)
Additionally, animating many animal tracks can be computationally intensive as well as create a potentially confusing plot, so for this lesson we will only be plotting one fish. We well subset our data by filtering where the animal_id is equal to NSBS-1393342-2021-08-10.
one_fish <- plot_data[plot_data$animal_id == "NSBS-1393342-2021-08-10",]
Preparing a Static Plot
Now that we have our data we can begin to create our plot. We will start with creating a static plot and then once happy with that, we will animate it.
The first thing we will do for our plot is download the basemap. This will provide the background for our plot. To do this we will use the get_stadiamap function from ggmap. This function gets a Stamen Map based off a bounding box that we provide. “Stamen” is the name of the service that provides the map tiles, but it was recently bought by Stadia, so the name of the function has changed. To create the bounding box we will pass a vector of four values to the argument bbox ; those four values represent the left, bottom, right, and top boundaries of the map.
To determine which values are needed we will use the min and max function on the mean_longitude and mean_latitude columns of our one_fish variable. min(one_fish$mean_longitude) will be our left-most bound, min(one_fish$mean_latitude) will be our bottom bound, max(one_fish$mean_longitude) will be our right-most bound, and max(one_fish$mean_latitude) will be our top bound. This gives most of what we need for our basemap but we can further customize our plot with maptype which will change what type of map we use, crop which will crop raw map tiles to the specified bounding box, and zoom which will adjust the zoom level of the map.
A note on maptype
The different values you can put for maptype: “terrain”, “terrain-background”, “terrain-labels”, “terrain-lines”, “toner”, “toner-2010”, “toner-2011”, “toner-background”, “toner-hybrid”, “toner-labels”, “toner-lines”, “toner-lite”, “watercolor”
basemap <-
get_stadiamap(
bbox = c(left = min(one_fish$mean_longitude),
bottom = min(one_fish$mean_latitude),
right = max(one_fish$mean_longitude),
top = max(one_fish$mean_latitude)),
maptype = "stamen_toner_lite",
crop = FALSE,
zoom = 7)
ggmap(basemap)
Now that we have our basemap ready we can create our static plot. We will store our plot in a variable called otn.plot so we can access it later on.
To start our plot we will call the ggmap function and pass it our basemap as an argument. To make our detection locations we will then call geom_point, supplying one_fish as the data argument. For the aesthetic we will make the x argument equal to mean_longitude and the y argument will be mean_latitude. This will orient our map and data properly.
We will then call geom_path to connect those detections supplying one_fish as the data argument. The aesthetic x will again be mean_longitude and y will be mean_latitude.
Lastly, we will use the labs function to add context to our plot including a title, a label for the x axis, and a label for the y axis. We are then ready to view our graph by calling ggplotly with otn.plot as the argument!
otn.plot <-
ggmap(basemap) +
geom_point(data = one_fish, aes(x = mean_longitude, y = mean_latitude), size = 2) +
geom_path(data = one_fish, aes(x = mean_longitude, y = mean_latitude)) +
labs(title = "NSBS Animation",
x = "Longitude", y = "Latitude", color = "Tag ID")
ggplotly(otn.plot)
Animating our Static Plot
Once we have a static plot we are happy with, we are ready for the final step of animating it! We will use the gganimate package for this, since it integrates nicely with ggmap.
To animate our plot we update our otn.plot variable by using it as our base, then add a label for the dates to go along with the animation. We then call transition_reveal, which is a function from gganimate that determines how to create the transitions for the animations. There are many transitions you can use for animations with gganimate but transition_reveal will calculate intermediary values between time observations. For our plot we will pass transition_reveal the first_detection information. We will finally use the functions shadow_mark with the arguments of past equal to TRUE and future equal to FALSE. This makes the animation continually show the previous data (a track) but not the future data yet to be seen (allowing it to be revealed as the animation progresses).
Finally, to see our new animation we call the animate function with otn.plot as the argument.
otn.plot <-
otn.plot +
labs(subtitle = 'Date: {format(frame_along, "%d %b %Y")}') +
transition_reveal(first_detection) +
shadow_mark(past = TRUE, future = FALSE)
animate(otn.plot)
Key Points
Animation with pathroutr
Overview
Teaching: 20 min
Exercises: 0 minQuestions
How can I create animal movement plots that avoid land?
Objectives
Basic animations using gganimate are great for many purposes but you will soon run into issues where your fish tracks are moving across land barriers, especially in more complex environments. This is because the geom used in the previous lesson choose the most direct path between two detection points. To avoid this we need to use a specific land avoidance tool. For our next animation we will use the pathroutr package which you can find out more about here.
We will begin in much the same way we did for our basic animation lesson with getting our data ready, but things will differ when we start to use pathroutr since there are specific data formats expected.
Preparing our Data
Just as in the basic animations lesson, we will only look at one fish. We will also filter down the data and only look at 5 detection events as an example subset due to the computational intensity of pathroutr and its calculations.
library(glatos)
library(sf)
library(gganimate)
library(tidyverse)
library(pathroutr)
library(ggspatial)
library(sp)
library(raster)
library(geodata)
detection_events <- #create detections event variable
read_otn_detections('nsbs_matched_detections_2022.csv') %>% # reading detections
false_detections(tf = 3600) %>% #find false detections
dplyr::filter(passed_filter != FALSE) %>%
detection_events(location_col = 'station', time_sep=3600)
plot_data <- detection_events %>%
dplyr::select(animal_id, mean_longitude,mean_latitude, first_detection)
one_fish <- plot_data[plot_data$animal_id == "NSBS-1393342-2021-08-10",]
There is one small tweak we are going to make that is not immediately intuitive, and which we’re only doing for the sake of this lesson. The blue sharks in our dataset have not given us many opportunities to demonstrate pathroutr’s awareness of coastlines. In order to give you a fuller demonstration of the package, we are going to cheat and shift the data 0.5 degrees to the west, which brings it more into contact with the Nova Scotian coastline and lets us show off pathroutr more completely. You do not need to do this with your real data.
one_fish_shifted <- one_fish %>% mutate(mean_longitude_shifted = mean_longitude-0.5)
Getting our Shapefile
The first big difference between our basic animation lesson and this lesson is that we will need a shapefile of the study area, so pathroutr can determine where the landmasses are located. To do this we will use the gadm function from the geodata library which gets administrative boundaries (i.e, borders) for anywhere in the world. The first argument we will pass to gadm is the name of the country we wish to get, in this case, Canada. We will specify level as 1, meaning we want our data to be subdivided at the first level after ‘country’ (in this case, provinces). 0 would get us a single shapefile of the entire country; 1 will get us individual shapefiles of each province. We must also provide a path for the downloaded shapes to be stored (./geodata here), and optionally a resolution. gadm only has two possible values for resolution: 1 for ‘high’ and 2 for ‘low’. We’ll use low resolution here because as we will see, for this plot it is good enough and will reduce the size of the data objects we download.
This is only one way to get a shapefile for our coastlines- you may find you prefer a different method. Regardless, this is the one we’ll use for now.
CAN<-gadm('CANADA', level=1, path="./geodata", resolution=2)
We only need one province, which we can select using the filtering methods common to R.
shape_file <- CAN[CAN$NAME_1 == 'Nova Scotia',]
This shapefile is a great start, but we need the format to be an sf multipolygon. To do that we will run the st_as_sf function on our shapefile. We also want to change the coordinate reference system (CRS) of the file to a projected coordinate system since we will be mapping this plot flat. To do that we will run st_transform and provide it the value 5070.
ns_polygon <- st_as_sf(single_poly) %>% st_transform(5070)
Formatting our Dataset
We will also need to make some changes to our detection data as well, in order to work with pathroutr. To start we will need to turn the path of our fish movements into a SpatialPoints format. To do that we will get the deploy_long and deploy_lat with dplyr::select and add them to a variable called path.
Using the SpatialPoints function we will pass our new path variable and CRS("+proj=longlat +datum=WGS84 +no_defs") for the proj4string argument. Just like for our shapefile we will need to turn our path into an sf object by using the st_as_sf function and change the CRS to a projected coordinate system because we will be mapping it flat.
path <- one_fish_shifted %>% dplyr::select(mean_longitude_shifted,mean_latitude)
path <- SpatialPoints(path, proj4string = CRS("+proj=longlat +datum=WGS84 +no_defs"))
path <- st_as_sf(path) %>% st_transform(5070)
We can do a quick plot to just check how things look at this stage and see if they are as expected.
ggplot() +
ggspatial::annotation_spatial(ns_polygon, fill = "cornsilk3", size = 0) +
geom_point(data = path, aes(x=unlist(map(geometry,1)), y=unlist(map(geometry,2)))) +
geom_path(data = path, aes(x=unlist(map(geometry,1)), y=unlist(map(geometry,2)))) +
theme_void()
Using pathroutr
Now that we have everything we need we can begin to use pathroutr. First, we will turn our path points into a linestring - this way we can use st_sample to sample points on our path.
plot_path <- path %>% summarise(do_union = FALSE) %>% st_cast('LINESTRING')
track_pts <- st_sample(plot_path, size = 10000, type = "regular")
The first pathroutr function we will use is prt_visgraph. This creates a visibility graph that connects all of the vertices for our shapefile with a Delaunay triangle mesh and removes any edges that cross land. You could think of this part as creating the viable routes an animal could swim through (marking the “water” as viable).
vis_graph <- prt_visgraph(ns_polygon, buffer = 100)
To reroute our paths around the landmasses we will call the prt_reroute function. Passing track_pts, md_polygon, and vis_graph as arguments. To have a fully updated path we can run the prt_update_points function, passing our new path track_pts_fix with our old path track_pts.
track_pts_fix <- prt_reroute(track_pts, ns_polygon, vis_graph, blend = TRUE)
track_pts_fix <- prt_update_points(track_pts_fix, track_pts)
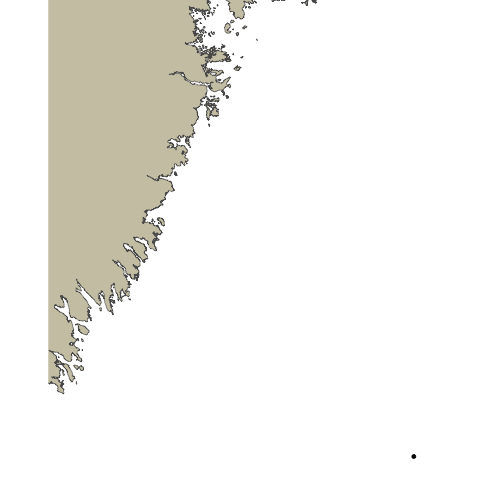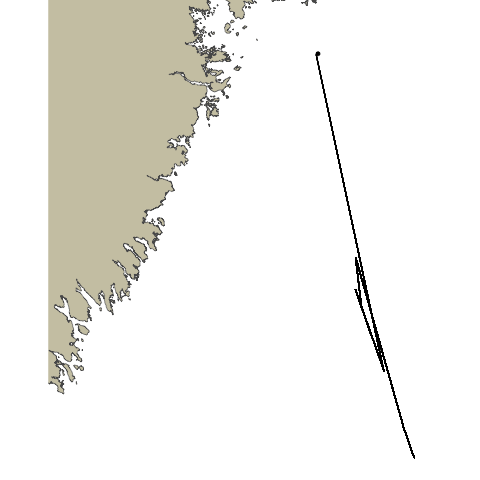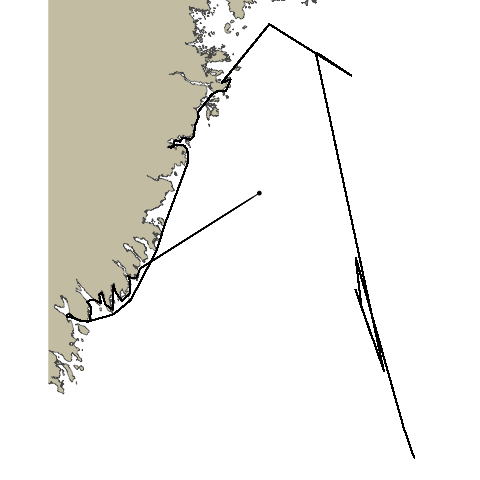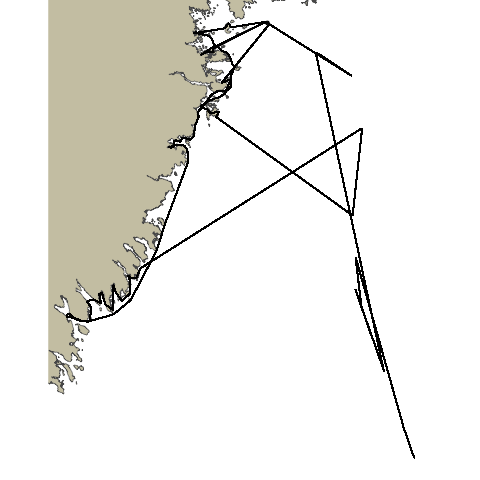
Now with our newly fixed path we can visualize it and see how it looks. We can also use this plot as the base plot for our animation.
For geom_point and geom_path we will pass in track_pts_fix for the data argument, but we will need to get a little creative for the x and y arguments in the aesthetic. track_pts_fix is a list of points so we will need a way to subset just the x and y values in order to supply them to the aesthetic. We will do this using map(geometry,1) to get a list of the values, and then unlist to turn that into a vector.
pathroutrplot <- ggplot() +
ggspatial::annotation_spatial(ns_polygon, fill = "cornsilk3", size = 0) +
geom_point(data = track_pts_fix, aes(x=unlist(map(geometry,1)), y=unlist(map(geometry,2)))) +
geom_path(data = track_pts_fix, aes(x=unlist(map(geometry,1)), y=unlist(map(geometry,2)))) +
theme_void()
pathroutrplot
Animating our New Path
With our plot in good order we are now able to animate! We will follow what we did in the basic animation lesson with updating our pathroutrplot variable by using it as the basemap, then adding extra information. Using the function transition_reveal and then shadow_mark, we will use the arguments of past equal to TRUE and future equal to FALSE. Then we are good to call the gganimate::animate function and watch our creation!
pathroutrplot.animation <-
pathroutrplot +
transition_reveal(fid) +
shadow_mark(past = TRUE, future = FALSE)
gganimate::animate(pathroutrplot.animation, nframes=100, detail=2)
A Note on Detail
You’ll note that the animation we’ve generated still crosses the landmass at certain points. This is a combination of several factors: our coastline polygon is not very high-res, our animation does not have many frames, and what frames it does have are not rendered in great detail. We can increase all of these and get a more accurate plot. For example:
- We can specify
resolution=1when downloading our shapefile from GADM.- We can increase the
nframesvariable in our call togganimate::animate.- We can pass
detail = 2or higher to the call togganimate::animate.All of these will give us an animation that more scrupulously respects the landmass, however, they will all bloat the runtime of the code significantly. This may not be a consideration when you create your own animations, but they do make it impractical for this workshop. Embedded below is an animation created with high-resolution polygons and animation parameters to show an example of the kind of animation we could create with more time and processing power.
Key Points
Spatial and Temporal Modelling with GAMs
Overview
Teaching: min
Exercises: 0 minQuestions
What are GAMs?
How can I use GAMs to visualise my data?
Objectives
GAM is short for ‘Generalized Additive Model’, a type of statistical model. In this lesson, we will be using GAMs to visualise our data.
While we are still developing this lesson as templated text like our other lessons, we can provide the Powerpoint slides for the GAMs talk given by Dr. Lennox at the CANSSI Early Career Researcher workshop. You can access the slides here.
The following code is meant to be run alongside this presentation.
require(BTN) # remotes::install_github("robertlennox/BTN")
require(tidyverse)
require(mgcv)
require(lubridate)
require(Thermimage)
require(lunar)
library(sf)
library(gratia)
theme_set<-theme_classic()+
theme(text=element_text(size=20), axis.text=element_text(colour="black"))+
theme(legend.position="top") # set a theme for ourselves
aur<-BTN::aurland %>%
st_transform(32633) %>%
slice(2) %>%
ggplot()+
geom_sf()
#Load RData
load("YOUR/PATH/TO/data/otn/troutdata.RDS")
# first plot of the data
troutdata %>%
ggplot(aes(dt, Data, colour=Data))+
geom_point()+
theme_classic()+
scale_colour_gradientn(colours=Thermimage::flirpal)
# mapping out the data
aur+
geom_point(data=troutdata %>%
group_by(lon, lat) %>%
dplyr::summarise(m=mean(Data)),
aes(lon, lat, colour=m))+
scale_colour_gradientn(colours=Thermimage::flirpal)+
theme(legend.position="top", legend.key.width=unit(3, "cm"))+
theme_bw()
# 4. going for smooth
a<-troutdata %>%
mutate(h=hour(dt)) %>%
bam(Data ~ s(h, bs="cc", k=5), data=., method="fREML", discrete=T)
b<-troutdata %>%
mutate(h=hour(dt)) %>%
bam(Data ~ s(h, bs="tp", k=5), data=., , method="fREML", discrete=T)
c<-troutdata %>%
mutate(h=hour(dt)) %>%
bam(Data ~ h, data=., , method="fREML", discrete=T)
tibble(h=c(0:23)) %>%
mutate(circular=predict.gam(a, newdata=.)) %>%
mutate(thin_plate=predict.gam(b, newdata=.)) %>%
# mutate(v3=predict.gam(c, newdata=.)) %>%
gather(key, value, -h) %>%
ggplot(aes(h, value, colour=key))+
geom_line(size=2)+
theme_set+
labs(x="Hour", y="Predicted Depth", colour="Model")+
scale_y_reverse(limits=c(20, 0))+
geom_hline(yintercept=0)+
coord_polar()
#6. model fitting vehicle
m1<-troutdata %>%
mutate(h=hour(dt)) %>%
mutate(foid=factor(oid)) %>%
gam(Data ~ s(h, k=5)+s(foid, bs="re"), data=., method="REML")
m2<-troutdata %>%
mutate(h=hour(dt)) %>%
mutate(foid=factor(oid)) %>%
bam(Data ~ s(h, k=5)+s(foid, bs="re"), data=., method="fREML", discrete=T)
tibble(h=c(0:23)) %>%
mutate(foid=1) %>%
mutate(gam=predict.gam(m1, newdata=., type="response", exclude=c("s(foid)"))) %>%
mutate(bam=predict.gam(m2, newdata=., type="response", exclude=c("s(foid)"))) %>%
mutate(i=gam-bam) %>%
gather(key, value, -h, -foid, -i) %>%
ggplot(aes(h, value, colour=i))+
geom_line(size=2)+
theme_set+
facet_wrap(~key)+
labs(x="Hour", y="Predicted temperature", colour="Difference between predictions")+
theme(legend.key.width=unit(3, "cm"))+
scale_colour_gradientn(colours=Thermimage::flirpal)
#8. check your knots
k1<-troutdata %>%
mutate(h=hour(dt)) %>%
bam(Data ~ s(h, bs="cc", k=5), data=., method="fREML", discrete=T)
k2<-troutdata %>%
mutate(h=hour(dt)) %>%
bam(Data ~ s(h, bs="cc", k=15), data=., , method="fREML", discrete=T)
tibble(h=c(0:23)) %>%
mutate("k=5"=predict.gam(k1, newdata=., type="response")) %>%
mutate("k=15"=predict.gam(k2, newdata=., type="response")) %>%
gather(key, value, -h) %>%
ggplot(aes(h, value/10, colour=key))+
geom_line(size=2)+
theme_set+
labs(y="Temperature", x="Hour", colour="model")
#9. temporal dependency
t1<-troutdata %>%
mutate(h=hour(dt), yd=yday(dt), foid=factor(oid)) %>%
group_by(foid, dti=round_date(dt, "1 hour")) %>%
dplyr::filter(dt==min(dt)) %>%
bam(Data ~ s(h, k=5, bs="cc")+s(yd, k=10)+s(foid, bs="re"), data=., method="fREML", discrete=T)
t2<-troutdata %>%
mutate(h=hour(dt), yd=yday(dt), foid=factor(oid)) %>%
group_by(foid, dti=round_date(dt, "1 hour")) %>%
bam(Data ~ s(h, k=5, bs="cc")+s(yd, k=10)+s(foid, bs="re"), data=., method="fREML", discrete=T)
expand_grid(h=c(12),
yd=c(32:60),
foid=1) %>%
mutate(partial_series=predict.gam(t1, newdata=., type="response", exclude=c("s(foid)"))) %>%
mutate(full_series=predict.gam(t2, newdata=., type="response", exclude=c("s(foid)"))) %>%
gather(key, value, -h, -foid, -yd) %>%
ggplot(aes(yd, value, colour=key))+
geom_point(data=troutdata %>%
mutate(h=hour(dt), yd=yday(dt), foid=factor(oid)),
aes(yday(dt), Data), inherit.aes=F)+
geom_path(size=2)+
theme_set+
labs(x="Date", y="Temperature", colour="Model")
# 10. spatial dependency
aur+
geom_point(data=troutdata %>%
group_by(lon, lat) %>%
dplyr::summarise(m=mean(Data)),
aes(lon, lat, colour=m))+
scale_colour_gradientn(colours=Thermimage::flirpal)+
theme(legend.position="top", legend.key.width=unit(3, "cm"))+
theme_bw()+
theme_set+
theme(legend.position="top", legend.key.width=unit(3, "cm"))+
labs(colour="mean temperature")
#11. interactions
mi<-troutdata %>%
mutate(h=hour(dt), yd=yday(dt), foid=factor(oid)) %>%
bam(Data ~ te(h, yd, bs=c("cc", "tp"), k=c(5, 10))+
s(foid, bs="re"), data=., family=Gamma(link="log"), method="fREML", discrete=T)
ms<-troutdata %>%
mutate(h=hour(dt), yd=yday(dt), foid=factor(oid)) %>%
bam(Data ~ s(h, bs="cc", k=5)+
s(yd, bs="tp", k=10)+
s(foid, bs="re"), data=., family=Gamma(link="log"), method="fREML", discrete=T)
p1<-expand_grid(h=c(0:23), yd=c(182:212)) %>%
mutate(foid=1) %>%
mutate(value=predict.gam(ms, newdata=., type="response", exclude=c("s(foid)"))) %>%
mutate(i="Simple model, AIC=426 801") %>%
ggplot(aes(yd, h, fill=value))+
geom_raster()+
scale_fill_viridis_c()+theme_set+
theme(legend.key.width=unit(3, "cm"))+
labs(x="Date", y="Hour", fill="Predicted temperature")+
facet_wrap(~i)
p2<-expand_grid(h=c(0:23), yd=c(182:212)) %>%
mutate(foid=1) %>%
mutate(value=predict.gam(mi, newdata=., type="response", exclude=c("s(foid)"))) %>%
mutate(i="Interaction model, AIC=425 805") %>%
ggplot(aes(yd, h, fill=value))+
geom_raster()+
scale_fill_viridis_c()+theme_set+
theme(legend.key.width=unit(3, "cm"))+
labs(x="Date", y="Hour", fill="Predicted temperature")+
facet_wrap(~i)
AIC(mi, ms)
gridExtra::grid.arrange(p1, p2)
# is it the moon?
a<-expand_grid(h=c(0:23), yd=c(182:212)) %>%
mutate(foid=1) %>%
mutate(value=predict.gam(ms, newdata=., type="response", exclude=c("s(foid)"))) %>%
mutate(i="Simple model, AIC= -262 975") %>%
mutate(dt=ymd("2022-12-31")+days(yd)) %>%
mutate(l=lunar::lunar.illumination(dt)) %>%
ggplot(aes(yd, h, fill=value))+
geom_raster()+
scale_fill_viridis_c()+theme_set+
theme(legend.key.width=unit(3, "cm"))+
labs(x="Date", y="Hour", fill="Predicted temperature")+
facet_wrap(~i)+
geom_point(data=expand_grid(h=c(0:23), yd=c(182:212)) %>%
mutate(foid=1) %>%
mutate(value=predict.gam(ms, newdata=., type="response", exclude=c("s(foid)"))) %>%
mutate(i="Simple model, AIC= -262 975") %>%
mutate(dt=ymd("2022-12-31")+days(yd)) %>%
mutate(l=lunar::lunar.illumination(dt)) %>%
distinct(dt, yd, l),
aes(yd, 10, size=l), inherit.aes=F, colour="white")
b<-expand_grid(h=c(0:23), yd=c(182:212)) %>%
mutate(foid=1) %>%
mutate(value=predict.gam(mi, newdata=., type="response", exclude=c("s(foid)"))) %>%
mutate(i="Interaction model, AIC=425 805") %>%
ggplot(aes(yd, h, fill=value))+
geom_raster()+
scale_fill_viridis_c()+theme_set+
theme(legend.key.width=unit(3, "cm"))+
labs(x="Date", y="Hour", fill="Predicted temperature")+
facet_wrap(~i)+
geom_point(data=expand_grid(h=c(0:23), yd=c(182:212)) %>%
mutate(foid=1) %>%
mutate(value=predict.gam(ms, newdata=., type="response", exclude=c("s(foid)"))) %>%
mutate(i="Simple model, AIC=426 801") %>%
mutate(dt=ymd("2022-12-31")+days(yd)) %>%
mutate(l=lunar::lunar.illumination(dt)) %>%
distinct(dt, yd, l),
aes(yd, 10, size=l), inherit.aes=F, colour="white")
gridExtra::grid.arrange(a, b)
########### the worked example
troutdata %>%
mutate(lun=lunar::lunar.illumination(dt)) %>%
ggplot(aes(lun, Data))+
geom_point()+
theme_set+
labs(x="Lunar illumination", y="Temperature")
troa<-troutdata %>%
mutate(foid=factor(oid)) %>%
mutate(lun=lunar::lunar.illumination(dt))
m0<-troa %>%
bam(Data ~ lun, data=., family=Gamma(link="log"))
tibble(lun=seq(0,1, by=0.1)) %>%
mutate(p=predict.gam(m0, newdata=., type="response")) %>%
ggplot(aes(lun, p))+
geom_point(data=troa, aes(lun, Data))+
geom_line(colour="red")+
theme_set+
labs(x="Moonlight", y="Temperature")
m0<-troa %>%
bam(Data ~ lun+s(foid, bs="re"), data=., family=Gamma(link="log"))
tibble(lun=seq(0,1, by=0.1)) %>%
mutate(foid=1) %>%
mutate(p=predict.gam(m0, newdata=., type="response", exclude=c("s(foid)"))) %>%
ggplot(aes(lun, p))+
geom_point(data=troa, aes(lun, Data, colour=factor(foid)))+
geom_line(colour="red")+
theme_set+
guides(colour=F)+
labs(x="Moonlight", y="Temperature")
m0<-troa %>%
bam(Data ~ s(lun, k=7)+s(foid, bs="re"), data=., family=Gamma(link="log"))
tibble(lun=seq(0,1, by=0.1)) %>%
mutate(foid=1) %>%
mutate(p=predict.gam(m0, newdata=., type="response", exclude=c("s(foid)"))) %>%
ggplot(aes(lun, p))+
geom_point(data=troa, aes(lun, Data, colour=factor(foid)))+
geom_line(colour="red")+
theme_set+
guides(colour=F)+
labs(x="Moonlight", y="Temperature")
m01<-troa %>%
mutate(yd=yday(dt), h=hour(dt)) %>%
bam(Data ~ s(lun, k=7)+s(foid, bs="re")+
s(h, bs="cc", k=5)+
s(lon, lat, k=15), data=., family=Gamma(link="log"))
BTN::aurland %>%
st_transform(32633) %>%
slice(2)
tibble(x=c(7.26, 7.32),
y=c(60.85, 60.87)) %>%
st_as_sf(., coords=c("x", "y")) %>%
st_set_crs(4326) %>%
st_transform(32633)
sp<-expand_grid(lon=seq(80077, 83583, by=10),
lat=seq(6770907, 6772740, by=10)) %>%
st_as_sf(., coords=c("lon", "lat")) %>%
st_set_crs(32633) %>%
st_intersection(BTN::aurland %>%
slice(1) %>%
st_transform(32633)) %>%
as(., "Spatial") %>%
as_tibble %>%
dplyr::rename(lon=coords.x1, lat=coords.x2) %>%
dplyr::select(lon, lat)
sp %>%
slice(1) %>%
expand_grid(., lun=seq(0, 1, by=0.1), h=c(0:23)) %>%
mutate(foid=1) %>%
mutate(value=predict.gam(m01, newdata=., exclude=c("s(foid)"))) %>%
ggplot(aes(h, lun, fill=value))+
geom_raster()+
scale_fill_gradientn(colours=Thermimage::flirpal)+
theme_classic()+
theme_set+
theme(legend.key.width=unit(3, "cm"))+
labs(x="Hour", y="Moonlight", fill="Predicted Temperature")
expand_grid(sp, lun=seq(0, 1, by=0.1), h=c(0, 12), yd=200) %>%
mutate(foid=1) %>%
mutate(p=predict.gam(m01, newdata=., type="response", exclude=c("s(foid)", "s(lon,lat)", "s(h)"))) %>%
ggplot(aes(lun, p))+
geom_point(data=troa, aes(lun, Data, colour=factor(foid)))+
geom_line()+
theme_set+
guides(colour=F)+
labs(x="Moonlight", y="Temperature")
sp %>%
expand_grid(., lun=seq(0, 1, by=0.3)) %>%
mutate(foid=1, h=1) %>%
mutate(value=predict.gam(m01, newdata=., type="response", exclude=c("s(foid)"))) %>%
ggplot(aes(lon, lat, fill=value))+
geom_raster()+
scale_fill_gradientn(colours=Thermimage::flirpal)+
theme_set+
theme(legend.key.width=unit(3, "cm"))+
labs(x="UTM (x)", y="UTM (y)", fill="Predicted Temperature")+
coord_fixed(ratio=1)+
facet_wrap(~lun)
gratia::draw(m01)
sp %>%
expand_grid(., lun=seq(0, 1, by=0.05), h=c(0:23)) %>%
mutate(foid=1) %>%
mutate(value=predict.gam(m01, newdata=., type="response", exclude=c("s(foid)", "s(lon,lat)"))) %>%
ggplot(aes(lun, value, colour=h))+
geom_point()+
theme_set+
theme(legend.key.width=unit(3, "cm"))+
scale_fill_gradientn(colours=Thermimage::flirpal)
coef(m01) %>%
data.frame %>%
rownames_to_column() %>%
as_tibble %>%
dplyr::filter(grepl("foid", rowname)) %>%
bind_cols(troa %>% distinct(foid)) %>%
left_join(troutdata %>%
group_by(oid) %>%
dplyr::summarise(m=mean(Data)) %>%
dplyr::rename(foid=oid) %>%
mutate(foid=factor(foid))) %>%
dplyr::rename(value=2) %>%
ggplot(aes(reorder(foid, value), value, colour=m))+
geom_point()+
coord_flip()+
theme_set+
labs(x="ID", y="Random intercept of temperature", size="Length (mm)", colour="True mean temperature")+
scale_colour_viridis_c()
Key Points
Introduction to the miscYAPS package
Overview
Teaching: 0 min
Exercises: 0 minQuestions
What is YAPS?
For what applications is YAPS well-suited?
How can I use the miscYAPS package to make working with YAPS easier?
Objectives
YAPS (short for ‘Yet Another Position Solver’) is a package originally presented in a 2017 paper by Baktoft, Gjelland, Økland & Thygesen. YAPS represents a way of positioning aquatic animals using statistical analysis (specifically, maximum likelihood analysis) in conjunction with time of arrival data, movement models, and state-space models. Likewise, miscYaps() is a package developed by Dr. Robert Lennox to provide more intuitive wrappers for YAPS functions, to ease the analysis process.
While we are still developing this lesson as templated text, as with the other lessons, we can provide the Powerpoint slides for the miscYaps talk given by Dr. Lennox at the CANSSI Early Career Researcher workshop. You can access the slides here.
The following code is meant to be run alongside this powerpoint.
remotes::install_github("robertlennox/miscYAPS")
require(yaps)
require(miscYAPS)
remotes::install_github("robertlennox/BTN")
require(BTN)
require(tidyverse)
require(lubridate)
require(data.table)
data(boats)
dets<-boats %>% pluck(1)
hydros<-dets %>%
dplyr::distinct(serial, x, y, sync_tag=sync) %>%
mutate(idx=c(1:nrow(.)), z=1)
detections<-dets %>%
dplyr::filter(Spp=="Sync") %>%
dplyr::select(ts, epo, frac, serial, tag)
ss_data<-boats %>% pluck(2) %>%
dplyr::rename(ts=dt) %>%
setDT
############
require(miscYAPS)
sync_model<-sync(hydros,
detections,
ss_data,
keep_rate=0.5,
HOW_THIN=100,
ss_data_what="data",
exclude_self_detections=T,
fixed=NULL)
plotSyncModelResids(sync_model)
sync_model<-sync(hydros,
detections,
ss_data,
keep_rate=0.5,
HOW_THIN=100,
ss_data_what="data",
exclude_self_detections=T,
fixed=c(1:9, 11:20))
fish_detections<-dets %>%
dplyr::filter(Spp!="Sync") %>%
mutate(tag=factor(tag)) %>%
dplyr::select(ts, epo, frac, tag, serial)
tr<-swim_yaps(fish_detections, runs=3, rbi_min=60, rbi_max=120)
data(aur)
btnstorel <- BTN::storel
raster::plot(btnstorel)
points(tr$x, tr$y, pch=1)
Key Points
Other OTN Telemetry Curriculums
Overview
Teaching: 0 min
Exercises: 0 minQuestions
How can I expand my learning?
Objectives
OTN has hosted other workshops in the past which contain different code sets that may be useful to explore after this workshop.
-
IdeasOTN Telemetry Workshop Series 2020: code available here and videos available on our YouTube here
Many of our Intro to R workshops are based upon this curriculum from The Carpentries.
Key Points